The parts of the lambda structure that you use for nonlinear constraints are lambda.ineqnonlin and lambda.eqnonlin.
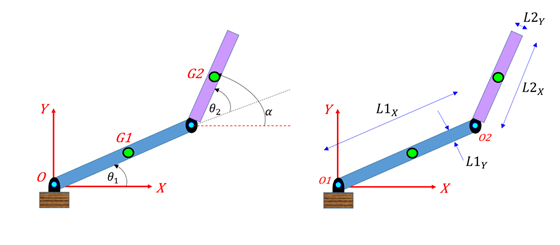
The Hessian function takes two input arguments: the position vector x, and the Lagrange multiplier structure lambda. Its Hessian is the Hessian of the Lagrangian see the User's Guide for more information.

This is because a nonlinearly constrained function needs to include those constraints in its Hessian. The interior-point algorithm requires its Hessian function to be written as a separate function, instead of being part of the objective function. Gradc = jacobian(c,x).' % transpose to put in correct formĬonstraint = matlabFunction(c,gradc, 'vars',) Since fmincon calls the objective function with column vectors, you must be careful to call matlabFunction with column vectors of symbolic variables.
#Matlab symbolic toolbox jacobian code
MatlabFunction generates code that depends on the orientation of input vectors. It is much more efficient to use matlabFunction. Therefore you should perform this calculation only once, and generate code, via matlabFunction, to call during execution of the solver.Įvaluating symbolic expressions with the subs function is time-consuming. This means that a symbolic gradient or Hessian has to be placed in the appropriate place in the objective or constraint function file or function handle.Ĭalculating gradients and Hessians symbolically can be time-consuming. Optimization gradients, and sometimes Hessians, are supposed to be calculated within the body of the objective or constraint functions. This requires you to translate between vectors and scalars. However, symbolic variables are scalar or complex-valued, not vector-valued. The results obtained yielded identical labeling patterns for metabolites as compared to the relaxation method.Optimization objective and constraint functions should be defined in terms of a vector, say x. The resulting equation system with the dimension of 546 x 546 was directly derived from 12 isotopomer balance equations. The method was applied to a network of lysine producing Corynebacterium glutamicum. This new method provided quick and robust convergence to the root of such equation systems in all cases tested. The solution of the complete system of equations is improved by implementing an iterative logical loop algorithm and using the analytical Jacobian matrix. These allow lumping of several consecutive isotopomer reactions into a single one.

The number of variables in the isotopomer distribution equation system is significantly reduced applying modified isotopomer mapping matrices.
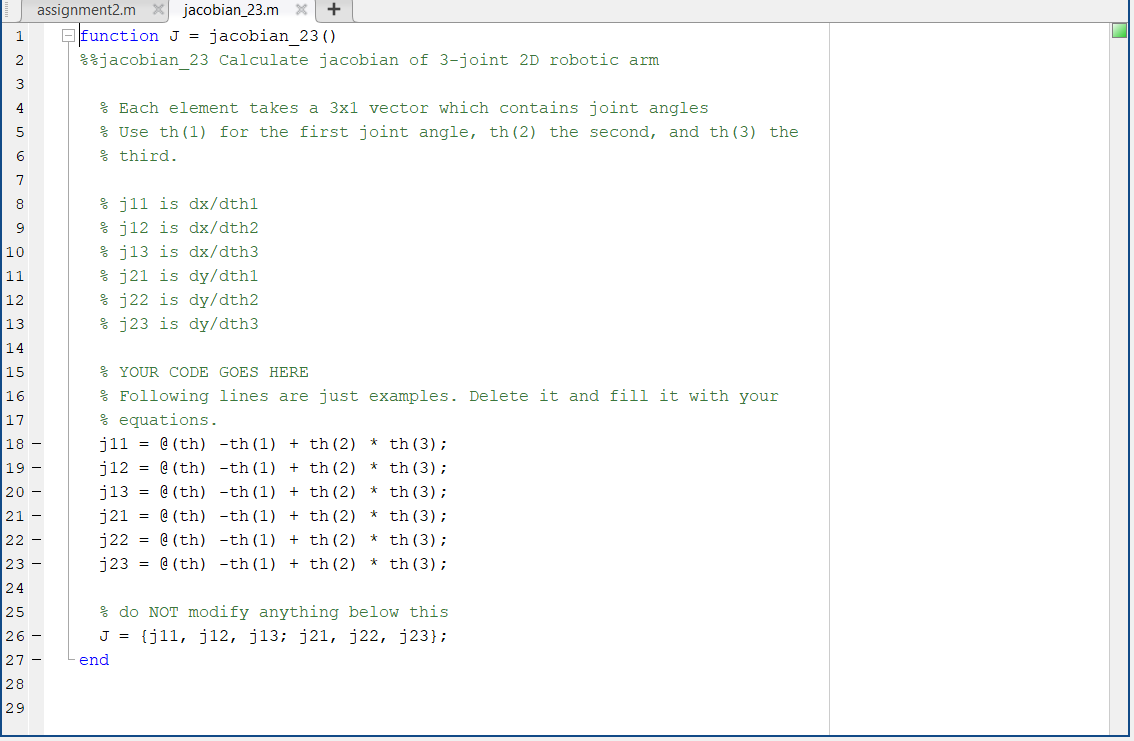
Using the symbolic toolbox of MATLAB the analytical solution of the stoichiometric balance equation system, isotopomer balances and the analytical Jacobian matrix of the total set of stoichiometric and isotopomer balances are created automatically. The latter are used to automatically calculate isotopomer mapping matrices. The only inputs required are the set of stoichiometric balances and the atom mapping matrices of all components of the reaction network. A novel method to accomplish efficient numerical simulation of metabolic networks for flux analysis was developed.

